Project Summary
The repository keeps the runnable Neural ODE pipeline and its recorded outputs, while the branch history documents two other comparison points: the physics-informed 3D attempt and the persistence baseline. The practical conclusion is consistent across all of them: the learned models are functional, but the tiny cohort is too small for them to surpass a simple last-scan predictor.
Main Code
The active pipeline is
scripts/run_neural_ode_pipeline.py. It supports full
prefix history, sliding windows, strict future holdout, and
separate-per-patient runs.
Data Context
The current local working set only contains
patient_007 and patient_067. The
larger LUMIERE cohort is the right next step for any claim about
generalization.
Branch Timeline
The work evolved from a physics-informed experiment into a cleaned Neural ODE pipeline, then into a prefix-history formulation that made the forecasting objective explicit.
Initial Physics Branch
A runnable 3D physics-informed attempt that acted as a historical comparison point, but did not beat the persistence baseline.
Cleaned Neural ODE Branch
The recovered Neural ODE idea was turned into a reproducible pipeline with strict future-week holdout and baseline evaluation.
Prefix-History Refinement
The sequence-to-one prefix formulation clarified the task and produced stable forecasts, but still lagged behind persistence.
Slim Main Branch
The current branch keeps the runnable pipeline, result artifacts, and a compact GitHub Pages summary.
Final Comparison
The key takeaway is that the model runs end to end, but the last-scan baseline remains the strongest method on this small local cohort.
| Method | Patient | Held-out week | MSE | Baseline MSE | MAE | Baseline MAE |
|---|---|---|---|---|---|---|
| Neural ODE strict holdout | patient_007 | 105 | 0.04119 | 0.00399 | 0.16927 | 0.03153 |
| Neural ODE strict holdout | patient_067 | 152 | 0.02426 | 0.00752 | 0.14075 | 0.04469 |
| Persistence baseline | patient_007 | 105 | 0.00399 | 0.00399 | 0.03153 | 0.03153 |
| Persistence baseline | patient_067 | 152 | 0.00752 | 0.00752 | 0.04469 | 0.04469 |
The learned model improves the forecasting setup, but not the final metric outcome on this cohort.
Results Gallery
The existing smoke-run outputs are kept in the repository and shown here directly so the page reflects actual experiments rather than only summary text.
Patient 007
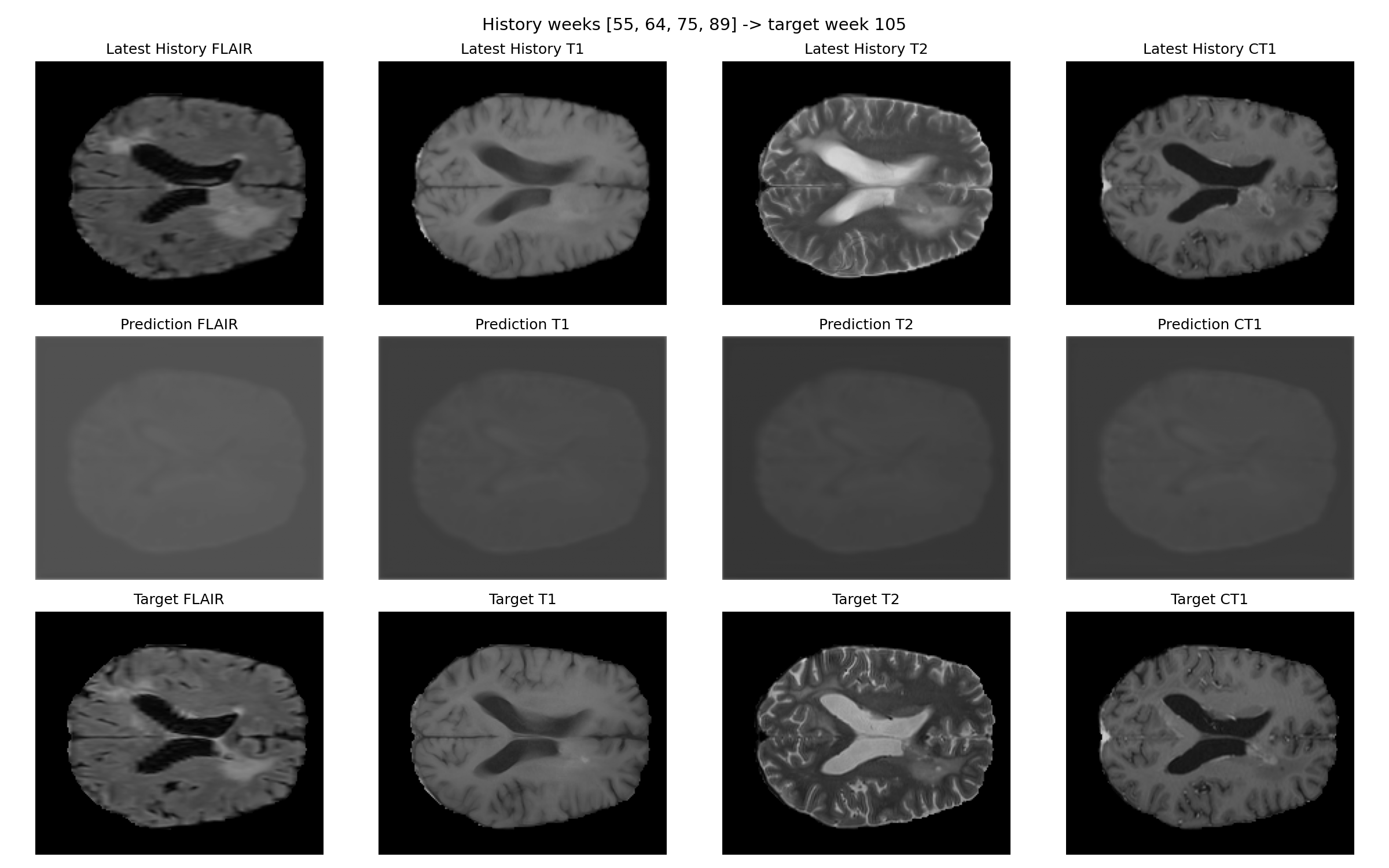
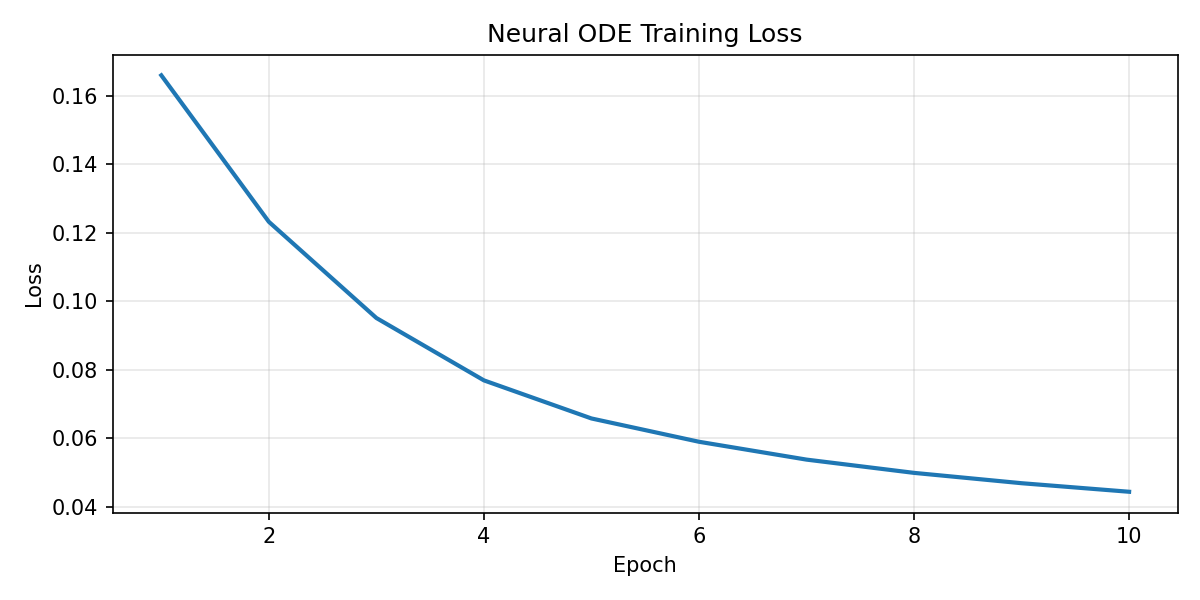
Patient 067
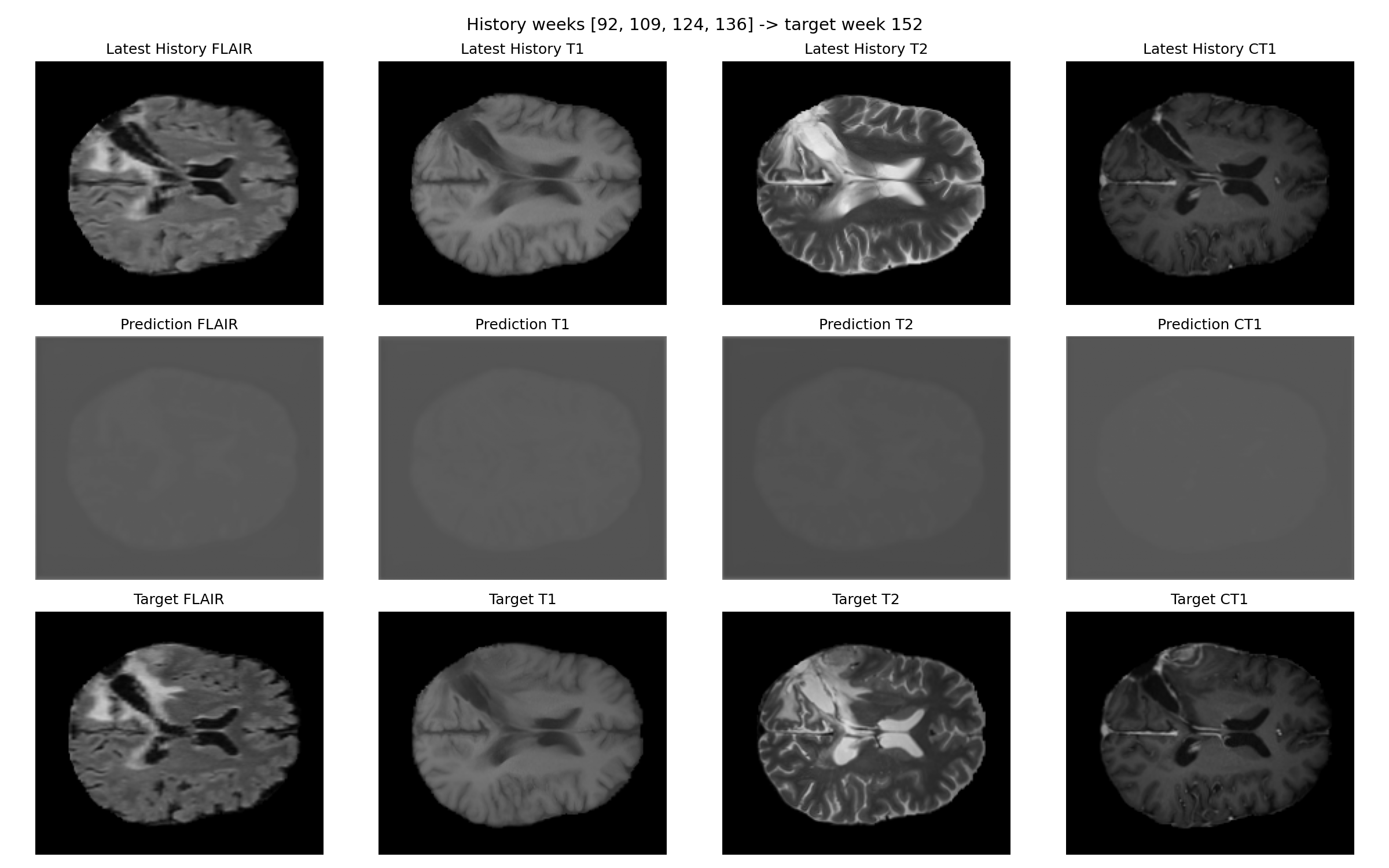
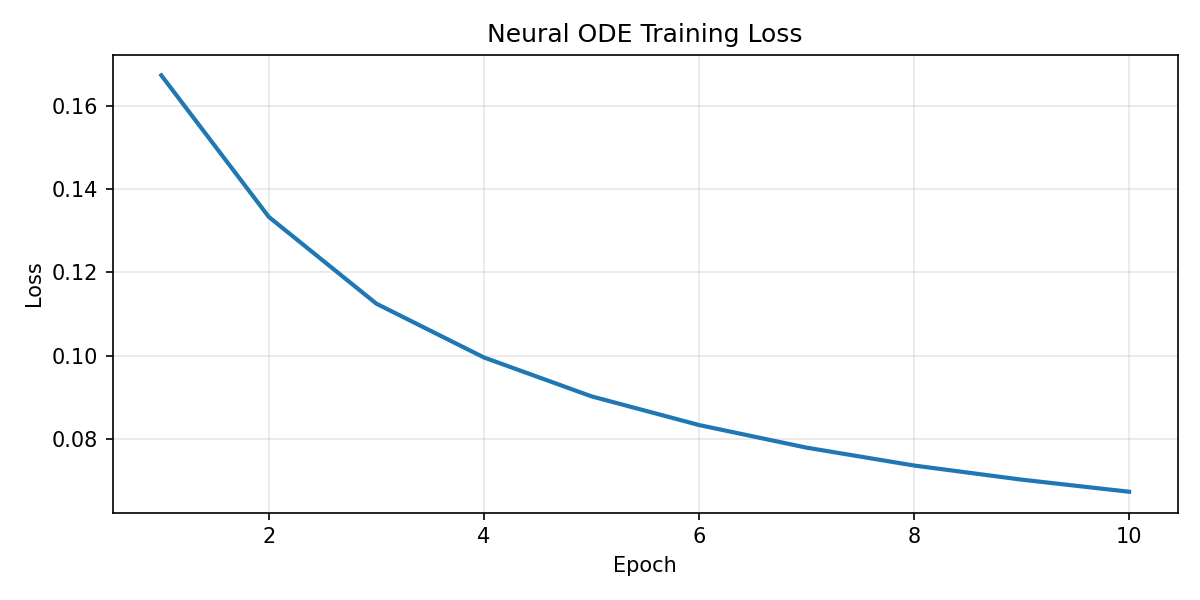
Interpretation
The physics-informed branch was a useful comparison point, but the practical non-Neural-ODE baseline is the persistence predictor. That simple approach still beats the learned methods here, which is why the repo is best read as a feasibility record rather than a final predictive claim.
What the learned models show
They demonstrate that the history-conditioned forecasting pipeline is implementable and that the models can produce plausible outputs for all four modalities.
What the baseline shows
The latest observed scan is already a very strong predictor on this small cohort, so any new model has to clear a high bar just to look useful.